-
Notifications
You must be signed in to change notification settings - Fork 6
Decision Tree in R : Metaprotein
Rahul Mondal edited this page Apr 21, 2021
·
17 revisions
We have implemented the following algorithms of Decision Trees for comparison of accuracy
- CART
- C 4.5
- C 5.0
Metaproteins as rows and, patient data of three types with samples from each being tested for the presence of metaproteins in columns (along with metaprotein demographics)
To suit our decision tree model, we removed the demographic columns for the dataset and transposed the data frame to turn metaproteins into columns/variables & patients as rows.
We created a class label "Patient type" which has 3 factors - C, UC & CD
We have taken half (1/2) of our Metaprotein Dataset to be used as Training Dataset & (1/2) to be used as Testing Dataset
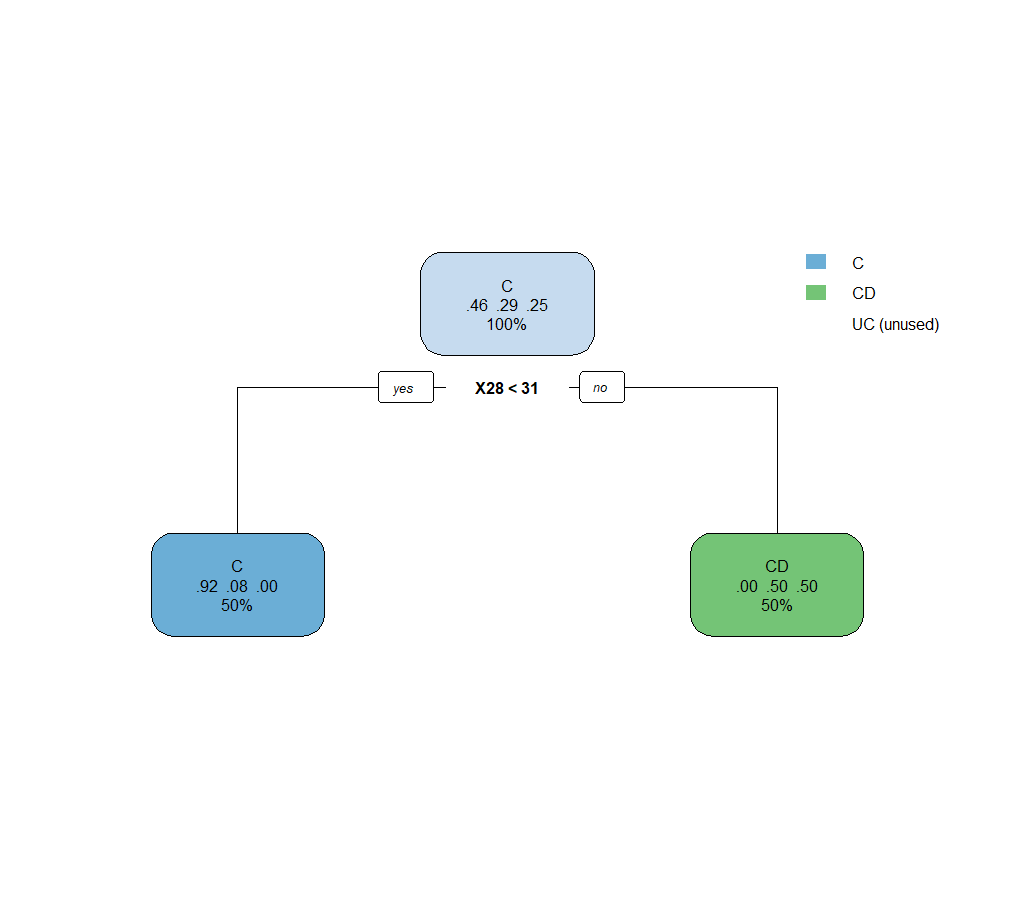
Accuracy = 15/24 = 62.5 %
Accuracy = 17/24 = 70.8 %
Accuracy = 17/24 = 70.8 %